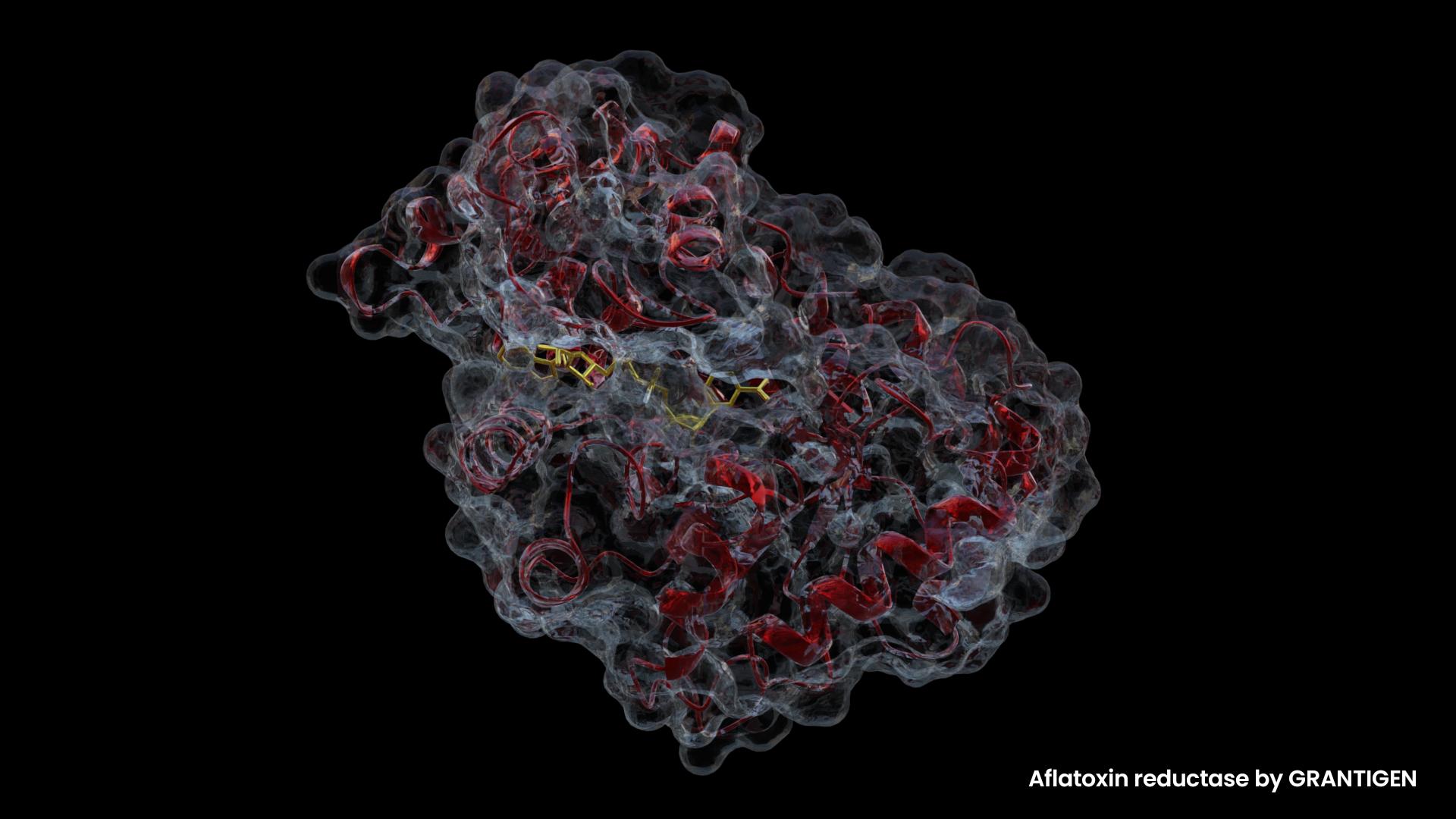
What are Aflatoxins?
Aflatoxins are metabolite products of Aspergillus, a genus of fungi frequently present in grain foods such as wheat, corn, rice, as well as peppers, ground nuts, tree nuts, and hay. Hay, of course, is not our food, but it can affect us indirectly.
There are over 400 micotoxins, of which Aflatoxin B1 (AFB1), one of the 21 aflatoxins, is the most harmful (1).
It is also the most important due to its deleterious effects on both our health and the food industry’s profit. Unfortunately, aflatoxins are thermally stable. Cooking does not degrade them (2).
ABF1 causes hepatotoxicity, nephrotoxicity, neurotoxicity, immunosuppression and adversely affects reproduction in laboratory animals. Most people have heard of aflatoxin in milk. The milk aflatoxin is designated as AFM1, and it is the hydroxylated version of AFB1 that is produced by dairy cows who eat contaminated feed (3). Both are classified as carcinogens, with M1 being slightly less toxic.
What is Aflatoxins’ mechanism of action?
The mechanism of action is complicated because the toxin affects many cellular pathways. As AFB1 gets processed in the liver, it gets oxidized by the P450 family of enzymes, forming a reactive epoxide. The epoxide readily binds DNA causing mutations (4).
Nothing that changes DNA is good for health. However, we were all taught that random mutations in DNA lead to gain of function and ultimately transition from species to species. How else would one explain the evolution of humans from monkeys unless, at some point in distant history, a random mutation produced an “ugly monkey” with a higher brain capacity, less fur, or the ability to walk upright, talk, or all of the above? That weird monkey must have happened by chance-say, one in a million. It’s safe to assume that human’ ancestors didn’t care about diversity so the poor monkey was probably ridiculed and unable to find a mate among the regular monkeys. Luckily, it must have found another “one in a million monkey” of the opposite sex within walking distance and produced offspring that built the foundation for our species.
We don’t know what triggered this serendipitous transformation in monkey DNA, especially what caused two pairs of their chromosomes to merge into one, as we have one pair of chromosomes less. Guessing that aflatoxins played a role in this is a guess as good as any other. These compounds are potent DNA modification agents, and DNA change is needed for the transformation of species. Maybe aflatoxins are not that bad after all? Maybe we owe them our existence? Shakespeare wrote, “There is nothing either good or bad, but thinking makes it so.” This perfectly aligns with evolution. Evolution knows neither good nor bad; it just happens. Therefore, we may not like dying of cancer, but it’s not relevant. Isn’t it interesting how painful it can be to observe one’s world from a non-anthropocentric point of view?
What is relevant, though, is that random mutations happen, and while most of the time they cause death and disability, occasionally they produce something new and something good. At least there is scientific consensus on that, regardless of the fact that we can’t see it happening in real life. Crooked-beaked finches and dark moths don’t really count, as those examples are weak.
After this detour into the possible mechanism of monkey-to-human transformation based on solid science doped with aflatoxins’ speculative role in evolution, we can now continue listing additional ways in which aflatoxins can harm us.
The toxin also depletes glutathione. Since glutathione is responsible for detoxifying the body, AFB1 can bind glutathione, causing elevated levels of reactive oxygen species due to the lack of this important endogenously produced antioxidant. For those who have wondered why it’s dangerous to take more Tylenol than what is written on the drug leaflet, the answer is the same. Tylenol too sequesters the slow-producing glutathione, leading to oxidative stress in mitochondria and ultimately triggering apoptosis (5).
Together with these two potential primary means of action, ABF1 affects steroid hormone biosynthesis, rethionoid and lipid metabolism, as well as AMPK, cAMP, insulin, mTOR, VEGF, FoxO, MAPK, TNF signaling pathways, just to name a few. The outcomes of large aflatoxin consumption could be liver cancer, lung cancer, and gastrointestinal cancer. The full list of pathways affected by aflatoxins is much longer. Readers can find out more from this review (6).
How is it possible that one molecule or one set of closely related molecules can affect so many biological processes? It seems that ABF1 is a Swiss army knife, but in a sinister way. It must be because it looks like something normally found in metabolism. Obviously, it shares an (evolutionary) design with steroid hormones and vitamin D. This could explain its interference with so many pathways. The full explanation will hopefully come in the near future because, although the author of this post cares about future generations, he cares a bit more about his health right now.
Another possible structure/function similarity interplay might be inferred from the fact that AFB1 downregulates the expression of the vitamin D receptor in the human osteasacroma cell line SAOS-2. Maybe that is why rickets is possible in places like Africa with abundant sunshine throughout the year, assuming an adequate diet is available (7)?
Let us now focus on the practical problems, such as:
1. Is it possible to remove aflatoxins from food?
2. How can we find out whether we have accumulated aflatoxins in our bodies?
3. Is it possible to detoxify from them?
1. Is it possible to remove aflatoxins from food?
It is, but with various degrees of success depending on the method (8). The first method is absorption on various types of materials such as charcoal. It is a physical method of low efficiency. In order to improve adsorption, the current adsorbents need to be improved. There have been continued efforts in this direction.
The second method is the chemical treatment of food. It’s more aggressive and efficient, but treating food with acids, bases, and oxidative agents changes the food as well, not just the toxin. Therefore, this method is not ideal either. Sodium metabisulphite and hydrogen peroxide combination seems to work quite well (9).
The third method relies on “good” microorganisms that can perform the detoxification of food (10). Good microorganisms can also absorb aflatoxins and degrade them. Food production processes may benefit from using good microorganisms, but at the same time, it may not be feasible to grow them on food just to prevent potential contamination with Aspergillus. Still, the microbial method is better than the previous two.
2. How can we find out whether we have accumulated aflatoxins in our bodies?
It is possible to detect accumulated toxins through the detection of their metabolites in urine. As we have already mentioned, AFM1 is excreted in milk, so it can be analyzed there. Blood can be analyzed as well, with various degrees of success due to the sensitivity of the method used (11).
3. Is it possible to detoxify our bodies?
Apparently yes, but the protocol has not yet been established. Curcumin has emerged as a potential scavenger, or better said, a protective agent for AFB1 (12). Curcumin has been investigated for many positive effects on health through its antibacterial, antiviral, and antitumor properties. The health benefits of this molecule go beyond what is listed here.
So far, curcumin’s protective actions have been investigated in vitro using BHF12 and HUC-PC cell lines, mice, rats, tilapia fish, chickens, and ducklings of different ages. In each case, it was determined that the group of animals ingesting curcumin in their diet contaminated with AFB1 had less tissue damage than the control group. Readers can find out more details in (12).
Grantigen needs to note that the experimental conditions used in the research mentioned do not closely mimic a real-life scenario because the amounts of both curcumin and AFB1 used were much larger than what would be the case in a real-life scenario. Even the ratio of the two may not necessarily be proportionately higher than what could happen in a regular diet in order to say that the experimental design just augmented the real-life scenario.
For example, in one study, ducklings were fed 400 mg/kg of curcumin per day. For a human of 70kg, that would be 28g of curcumin per day. Since it doesn’t come in pure form but rather in turmeric powder that contains about 3% of the active component, the amount of turmeric powder would end up being 933 g per day! This is the high end of the experiment design range. According to the duckling massacre paper, the protective effects of curcumin were confirmed in the tissues of 21-day-old ducklings (12). Apart from its macabre nature, this is a really good review that we recommend to anyone who wants to learn more on the topic but doesn’t have time to sift through the mountain of literature on aflatoxins.
In some other experiments, chickens were fed with the amount of turmeric that would be equivalent to 31 g of turmeric per day, which for an average human is possible. Considering the other beneficial effects of curcumin, this amount of turmeric ingested per day could potentially be a good addition to the western diet. Can it go much lower and still work?
Despite all of this, the best way to protect oneself from aflatoxins is to avoid them. It is more or less possible, but it also greatly depends on the region of the world. For example, hot and humid places are more likely to have elevated amounts of toxins in food, especially if this is coupled with lax food regulations in that country. In contrast, cold and arid countries that are regulated “to the bone,” like northern Europe and Canada, are less likely to have the toxin lurking in food, as that is one of the things that is checked before food is brought to consumers. Milk is easy to avoid because everyone who is not an infant can easily live without it. Yogurt contains the good microbes that are Aspergillus’ natural enemies; therefore, it could be safer than milk. Grains are everywhere, so they are difficult to avoid, and for that reason, one would end up relying on the government to protect him, which is always a scary thought.
Let us finish on a positive note:
So what happens if we consume low amounts of aflatoxin over a long period of time? The amounts that are not going to produce acute poisoning According to the study on 131 patients conducted in Italy, where in certain regions contamination is known to be present, but at low levels, 81% of hepatocellular carcinoma patients did not have traces of AFB1 in the liver cells (13). Of the remaining 19% who had traces of AFB1, only a few had high levels of the toxin in the cells. This study does not support a link between hepatocellular carcinoma and low ABF1 levels. Where was the negative control in this study anyway? Wouldn’t a negative control composed of healthy people who could potentially have mixed results in regards to traces of ABF1 in liver cells completely destroy the fear of low doses of aflatoxins? This may not be a good study to publish for a lab that gets funded to study aflatoxins, hence the lack of negative control.
Another study found aflatoxin present in the sera of 64% of primary liver cancer patients, while at the same time finding hepatitis B in 69% of the patients (14). This shows a strong correlation between the two, but the study doesn’t tell us what the correlation would be if there was no hepatitis B present. The study was conducted in Sub-Saharan Africa, a region that is at high risk of food contamination by Aspergillus; therefore, the comparison with the European study is not easy, or maybe not possible at all. We may have missed the paper we were looking for, but to Grantigen, it seems that the definitive link between low doses of aflatoxin exposure and hepatocellular carcinoma in the absence of other risk factors has not been established.
References:
1. Bennett JW, Klich M. Mycotoxins. Clin Microbiol Rev. 2003 Jul;16(3):497-516. doi: 10.1128/CMR.16.3.497-516.2003. PMID: 12857779; PMCID: PMC164220.
2. Yamada M, Hatsuta K, Niikawa M, Imaishi H. Detoxification of Aflatoxin B1 Contaminated Maize Using Human CYP3A4. J Microbiol Biotechnol. 2020 Aug 28;30(8):1207-1213. doi: 10.4014/jmb.2003.03032. PMID: 32423188; PMCID: PMC9728267.
3. Dhakal A, Hashmi MF, Sbar E. Aflatoxin Toxicity. 2023 Feb 19. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2023 Jan–. PMID: 32491713.
4. Johnson WW, Guengerich FP. Reaction of aflatoxin B1 exo-8,9-epoxide with DNA: kinetic analysis of covalent binding and DNA-induced hydrolysis. Proc Natl Acad Sci U S A. 1997 Jun 10;94(12):6121-5. doi: 10.1073/pnas.94.12.6121. PMID: 9177180; PMCID: PMC21012.
5. Benkerroum N. Chronic and Acute Toxicities of Aflatoxins: Mechanisms of Action. Int J Environ Res Public Health. 2020 Jan 8;17(2):423. doi: 10.3390/ijerph17020423. PMID: 31936320; PMCID: PMC7013914.
6. Marchese S, Polo A, Ariano A, Velotto S, Costantini S, Severino L. Aflatoxin B1 and M1: Biological Properties and Their Involvement in Cancer Development. Toxins (Basel). 2018 May 24;10(6):214. doi: 10.3390/toxins10060214. PMID: 29794965; PMCID: PMC6024316.
7.Costanzo P, Santini A, Fattore L, Novellino E, Ritieni A. Toxicity of aflatoxin B1 towards the vitamin D receptor (VDR). Food Chem Toxicol. 2015 Feb;76:77-9. doi: 10.1016/j.fct.2014.11.025. Epub 2014 Dec 4. PMID: 25483621.
8. Guan Y, Chen J, Nepovimova E, Long M, Wu W, Kuca K. Aflatoxin Detoxification Using Microorganisms and Enzymes. Toxins (Basel). 2021 Jan 9;13(1):46. doi: 10.3390/toxins13010046. PMID: 33435382; PMCID: PMC7827145.
9. Karlovsky P, Suman M, Berthiller F, De Meester J, Eisenbrand G, Perrin I, Oswald IP, Speijers G, Chiodini A, Recker T, Dussort P. Impact of food processing and detoxification treatments on mycotoxin contamination. Mycotoxin Res. 2016 Nov;32(4):179-205. doi: 10.1007/s12550-016-0257-7. Epub 2016 Aug 23. PMID: 27554261; PMCID: PMC5063913.
10. Yamada M, Hatsuta K, Niikawa M, Imaishi H. Detoxification of Aflatoxin B1 Contaminated Maize Using Human CYP3A4. J Microbiol Biotechnol. 2020 Aug 28;30(8):1207-1213. doi: 10.4014/jmb.2003.03032. PMID: 32423188; PMCID: PMC9728267.
11. Wild CP, Chapot B, Scherer E, Den Engelse L, Montesano R. Application of antibody methods to the detection of aflatoxin in human body fluids. IARC Sci Publ. 1988;(89):67-74. PMID: 3198233.
12. Dai C, Tian E, Hao Z, Tang S, Wang Z, Sharma G, Jiang H, Shen J. Aflatoxin B1 Toxicity and Protective Effects of Curcumin: Molecular Mechanisms and Clinical Implications. Antioxidants (Basel). 2022 Oct 14;11(10):2031. doi: 10.3390/antiox11102031. PMID: 36290754; PMCID: PMC9598162.
13. Gramantieri L, Gnudi F, Vasuri F, Mandrioli D, Fornari F, Tovoli F, Suzzi F, Vornoli A, D’Errico A, Piscaglia F, Giovannini C. Aflatoxin B1 DNA-Adducts in Hepatocellular Carcinoma from a Low Exposure Area. Nutrients. 2022 Apr 15;14(8):1652. doi: 10.3390/nu14081652. PMID: 35458213; PMCID: PMC9024438.
14. Tchana AN, Moundipa PF, Tchouanguep FM. Aflatoxin contamination in food and body fluids in relation to malnutrition and cancer status in Cameroon. Int J Environ Res Public Health. 2010 Jan;7(1):178-88. doi: 10.3390/ijerph7010178. Epub 2010 Jan 18. PMID: 20195440; PMCID: PMC2819783. (63% in liver cancer)